The quest for a genetic basis for everything, with the exception of a very few and very bad diseases, has been disappointing – our genetic inheritance is polygenic, where hundreds or thousands of genes each make an exceedingly small contribution. Polygenic scores (PGS), the measurement that comes from genome-wide association studies (GWAS), summates those small contributions and has some, but not great predictive value.
Polygenic scores are not “portable.”
PGS can provide “informative predictions” for highly heritable qualities like height, at around 25% and not so much for less heritable phenotypes, like “educational attainment,” about 13%. One clear limitation to polygenic scores is racial differences, for example, the presence of an Asian genetic disposition to develop uncomfortable flushing when drinking alcohol. Polygenic scores derived for one racial group rarely maintain their predictive value for another, even when the phenotype is highly heritable. For example, using the UK Biobank’s PGS for height improves predictability by about 11% in UK subjects, but that improvement drops to 3% in the Japanese population.
Bottom line, PGS scores developed for one racial group are not transferable, portable is the author’s word choice, to another. Other words that come to mind are that polygenic scores are brittle or fragile.
Nurture and Nature
The underlying reason is that GWAS are not that precise in identifying “causal variants,” they recognize an aggregate of possibilities; and those possibilities are impacted by other factors, more specifically how frequent are the variants in the population and a more exotic term, linkage disequilibrium. DO NOT PANIC. It merely means that some genes are more likely inherited together rather than independently; it has to do with genetic groupings already present in the individuals creating the inheritance – the parents.
Choosing our mates involves a lot of factors, but one common thread is described as assortative mating (I promise this is the last new term) which means that we frequently choose mates with similar phenotypes, like height or education or income. And to the extent that our choices change from culture to culture, this may play a role in the linkage disequilibrium and the derived polygenic score. It may not be sufficient to say that a PGS was determined along racial lines; it may require more demographic or cultural stratification to give a PGS a substantial predictive value.
A new study in bioRxiv attempts to answer the question of how demographics does or does not alter the predictive value of PGS.
The Study
The researchers used the UK Biobank dataset restricting themselves to “White British,” about 338,000 individuals. They derived PGS for subgroups stratified them by age, the ratio of females to males, and socioeconomic status, using a randomly selected subset to determine a PGS and the rest of population as the test set. In each case, they found that stratification changed the PGS predictive value. For example, in predicting BMI, the PGS of a younger age cohort was 125% more predictive than the PGS derived for the oldest age cohort. The variations within this one racial grouping, “White British,” when polygenic scores were calculated by age, gender and socioeconomic status varied to the same degree the scores changed along racial lines.
It isn’t nature or nurture; it is a complex interaction of the two. Polygenic scores may be much less portable than we thought, we need to know not only the racial characteristics but several socioeconomic characteristics as well if we are to develop clinically useful polygenic scoring. Some of those characteristics are easy, smoking may well influence genetics and assortative mating, but other aspects will be Rumsfeldian unknown unknowns.
GWAS derived polygenic scores have begun to show us the effect of genetic on highly heritable traits, like eye color or height. But this paper demonstrates that for less heritable conditions, like years spent in school or smoking, that PGS quickly lose their predictive value when you fail to define the population being scored more precisely. The organizations, private and public, collecting our genetic data are already aware that they must expand their participants along racial lines, this study indicates that they must also take into account a host of other socio-demographics. It is early in our understanding and development of PGS, it is struggling to move from the laboratory to clinicians and policymakers, but it is not ready for primetime just yet; for most of us, it remains edutainment.
Source: Variable prediction accuracy of polygenic scores within an ancestry group bioRxiv preprint DOI:10.1101/629949
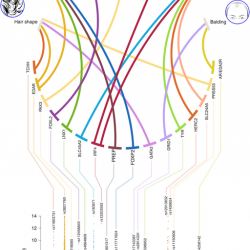
Images courtesy of the following individuals: Kaustubh Adhikari, Tania Fontanil, Santiago Cal, Javier Mendoza-Revilla, Macarena Fuentes-Guajardo, Juan-Camilo Chacón-Duque, Farah Al-Saadi,Jeanette A. Johansson, Mirsha Quinto-Sanchez, Victor Acuña-Alonzo, Claudia Jaramillo, William Arias, Rodrigo Barquera Lozano, Gastón Macín Pérez, Jorge Gómez-Valdés,Hugo Villamil-Ramírez, Tábita Hunemeier, Virginia Ramallo, Caio C. Silva de Cerqueira,Malena Hurtado,Valeria Villegas, Vanessa Granja, Carla Gallo, Giovanni Poletti, Lavinia Schuler-Faccini, Francisco M. Salzano,Maria-Cátira Bortolini, Samuel Canizales-Quinteros, Francisco Rothhammer, Gabriel Bedoya, Rolando Gonzalez-José, Denis Headon, Carlos López-Otín, Desmond J. Tobin,David Balding and Andrés Ruiz-Linares.