Last month there was talk of a 20 percent cut in funding for the National Institutes of Health, which is the primary source of academic life sciences grants. Prompted by science media concern, much of the public wondered about the potential impact on biomedical research. Pharmacologist David Kroll of Chemical & Engineering News expressed it well on Twitter:
"Combine the administration's harsh climate toward pharma profits with a proposed 20% NIH budget cut...where will new drugs come from?"
It’s a very common question. People who are not familiar with the drug discovery process often believe that government-funded research is the main driver of innovation. But this is false. Industry invents almost all drugs. When asked about what drugs are discovered in academia, Kroll noted that they don't do much but also expressed where academia does have importance (1).
"Virtually none are 'developed' by academia but 10-15% are discovered there...Academia contributes to revealing the pathophysiology for drug discovery and helps clarify mechanisms."
Despite the public belief that government-funded academia is the driver of drug development, they lack the billions of dollars, the work force, and the multidisciplinary expertise to overcome significant hurdles that stand between the lab and the pharmacy. Only the pharmaceutical industry can do this, and it is still plenty tough, even with billions of dollars and a decade or more of time to try.
But this does not mean that academia doesn't have its place in the discovery-development process. There is no better example of the synergy between the two than a breakthrough that is near and dear to me—hepatitis C. For ten years, I was stuck right in the middle of it. At that time, almost every major drug company and dozens of smaller ones were working on it. In the end—25 years after the virus was discovered—both industry and academia had played crucial, though different, roles which resulted in drugs that cure the disease.
First, some background. There are two basic types of assays (tests) that are used to determine whether any given molecule might be useful in treating a disease or infection. I'll use viruses as an example. One type of assay is enzyme-based. All viruses make enzymes, each of which performs a function to enable it to replicate. It is possible to isolate and grow the individual enzymes, and see if there any chemical compounds that might inhibit (block) them.
These assays are usually automated, which enables thousands (sometimes millions) (2) of chemical compounds to be tested to determine whether any of them inhibit the function of the enzyme, and how strongly. Presumably, a good inhibitor of an essential viral enzyme would then stop the sequence of events that are required for the virus to replicate (3).
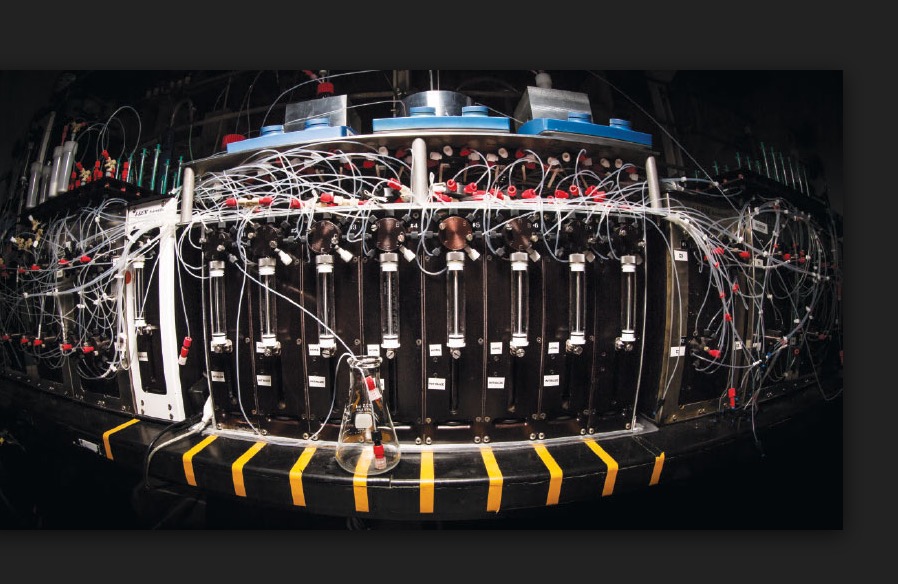

An automated chemical synthesis apparatus. Photo: M. Burke, University Of Illinois, Urbana-Champaign
I used the term "presumably" for a reason. It's not that simple. It is a very long trip from a compound that inhibits an enzyme in a test tube in your lab to the pharmacy. One of the many hurdles is especially frustrating— the inhibitor may work amazingly well in inhibiting a given enzyme in a test tube assay, but it will often lack the chemical and physical properties that enable it to get into cells, which is where the viral replication occurs. If the inhibitor cannot enter a cell it will not prevent replication (4). This lack of what we call cell permeability is the bane of drug discovery chemists, who often find themselves with a collection of highly potent inhibitors that are ultimately useless because they cannot cross cell membranes.
Fortunately, there is another approach—the cell-based assay. It is conceptually the reverse of the enzyme-based assay. Large collections of compounds are tested for their ability to inhibit a particular process that takes place within a cell, for example a virus replicating. If the compound inhibits replication in the cell this is great, but you do not know which enzyme it is inhibiting (the target). This must be determined later. But it is far easier to determine how something works inside a cell than to struggle to get a compound into the cell when it doesn't want to go there.
Simple enough then? No. Early hepatitis C research was crippled because there was no cell-based assay. Astoundingly, hepatitis C, which replicates like mad in your liver, will not do so in isolated liver cells. This is true for other viruses, like norovirus, also. No one really knows why.
So, in the absence of a cell-based assay, what to do? One possibility would be to select the best inhibitor(s) and test them in an animal model of the infection. Not only does this approach cause pharmaceutical employee madness (it has a very very low chance of success) the best animal model for hepatitis C is chimps, which are not used any more. So we hepatitis C researchers were operating with both hands tied behind our backs, a bag over our heads, and going nowhere fast.
That is, until academia came to the rescue.
With the use of molecular biology, Ralf Bartenschlager, (Heidelberg University in Germany) and Charles Rice (Washington University School of Medicine) invented the HCV subgenomic replicon. The science is very complex, but in short, the replicon is a modified virus in which the genes that make the structural components of the virus—the envelop and capsid— are absent.
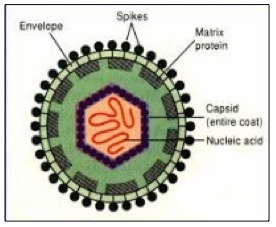
The structure of a typical virus. The envelope is the outer layer, which holds the antigens, The capsid contains the genetic material. Both are considered to be structural proteins.
In the replicon. the genes that make the capsid and envelope- the structural part of the virus - are removed. Only the genes that participate in the replication process remain. When the replicon is put into cultured liver cancer cells (5) it behaves (mostly) like the virus itself; it makes copies of itself, in other words, replicates. This gave chemists and virologists a tool to search for compounds that inhibited a surrogate of HCV.
Since the replicon is an artificial construct, there was initial concern that it might not be predictive of viral replication in living beings. But it is. The HCV replicon has been validated, and its use is now standard practice in hepatitis C. Sovaldi, the first legitimate direct-acting cure for hepatitis C, was selected (6) based on of its potency against the HCV replicon—a predictor of the drug's ability to stop viral replication at reasonable doses in people.
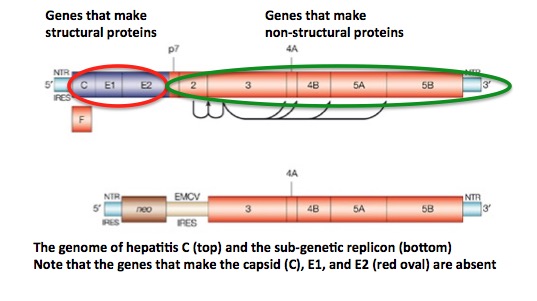
The hepatitis C subgenomic replicon. Image (modified): Nature.com
Without the sub-genomic replicon developed in universities, there was no way to predict whether a drug would stop hepatitis C. Without the thousands of analogs made in dozens of drug companies, Sovaldi never gets discovered at all.
A cure for one of the most important viral infections on earth (about 150 million people) resulted from drug companies getting key help from the research of academic institutions. There is a well-defined function for both and that is why in the rush to create smaller government and cheaper drugs, we should not penalize research in either arenas, especially if it will result in drugs like Sovaldi that help the entire world.
Notes:
(1) One surrogate measure of innovation is the name(s) on the patent. The only people who can appear on a patent are the inventors of a drug, medical device, etc. Even one extra (or missing) name can invalidate the patent. If you look at drug patents, most of them will contain the name(s) of scientists of drug companies. I am listed as an inventor on 25 of them. This means the company invented the drug, not a university or the NIH.
(2) Drug companies have libraries of compounds—hundreds of thousands, or even millions—of unique chemical compounds, which are used for high throughput screening. These libraries consists of chemicals that were synthesized for other programs, often many years ago. A small sample of every newly synthesized chemical compound is set aside for this. Alternatively, collections of compounds can be bought from specialty companies that make their own libraries, solely for the purpose of selling them for high throughput screening.
(3) The best example of the use of specific enzyme inhibition, which ultimately led to the discovery of vital drugs is HIV. There are approved drugs that target specific enzymes or receptors that inhibited specific enzymes that are essential for HIV replication that changed the infection from certain death sentence into a chronic, manageable disease.. The strategies employed for hepatitis C research were based largely on those that were discovered for HIV/AIDS.
(4) Viruses are obligate parasites. This means that in the absence of a host cell, replication cannot take place.
(5) Cell-based assays usually use a cancerous variant of the cell in question. This enable scientists to keep growing the cells (this is called immortality). The cells used for HCV research are called Huh-7, and are all derived from a sample taken from a liver tumor in Japanese man in 1982.




