Part of the global effort to discover new antibiotics involves inventing new techniques to analyze the ones we already have. The idea is that the more we learn about how antimicrobials work at the molecular level, the easier it will be to find or synthesize novel ones.
One way to learn about how antibiotics work is to visualize their accumulation within bacterial cells. But this is no easy feat. Several imaging techniques already exist, such as monitoring fluorescence or tracking molecules using radioactive labels, but these methods suffer from various drawbacks. So, a team of researchers from Penn State University and the pharmaceutical and biotechnology giant Novartis set about inventing a new technique.
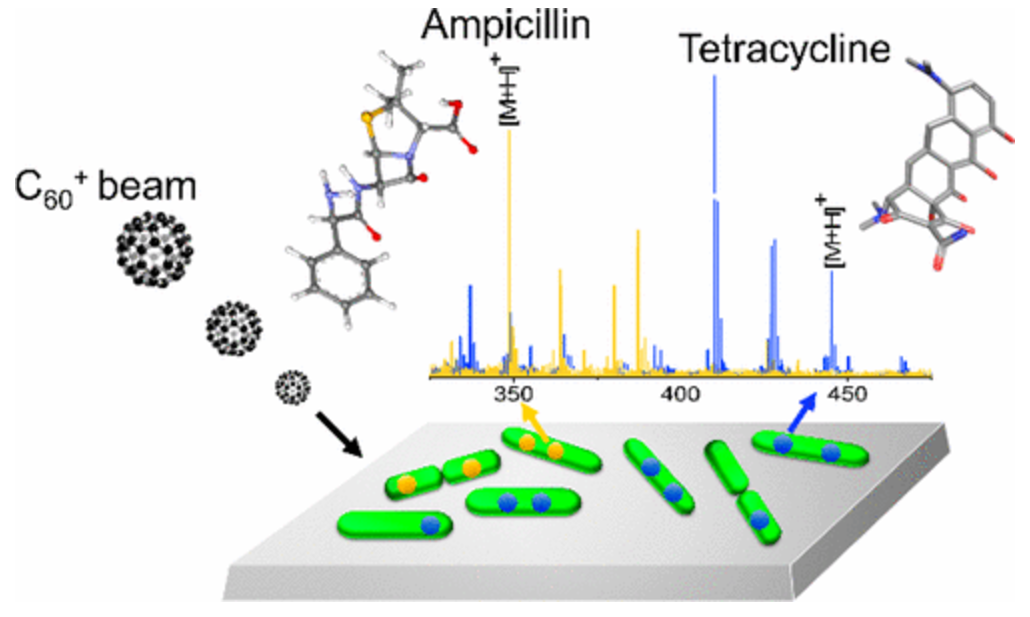 The researchers turned to an established protocol known as secondary ion mass spectrometry. This technique shoots a beam of ions (the "primary ions") at a sample, causing the molecules they hit to get blasted off (the "secondary ions"). These secondary ions are then detected by a mass spectrometer. (See image on right.)
The researchers turned to an established protocol known as secondary ion mass spectrometry. This technique shoots a beam of ions (the "primary ions") at a sample, causing the molecules they hit to get blasted off (the "secondary ions"). These secondary ions are then detected by a mass spectrometer. (See image on right.)
In this experiment, the authors applied this established methodology to a new purpose: To detect the localization of antibiotics inside bacteria. They aimed a beam of positively charged buckyballs at E. coli that had been treated with antibiotics. When the beam hits the bacteria, the cells slowly erode. (Think of how a paleontologist reveals a fossil as he brushes away dirt layer by layer.)
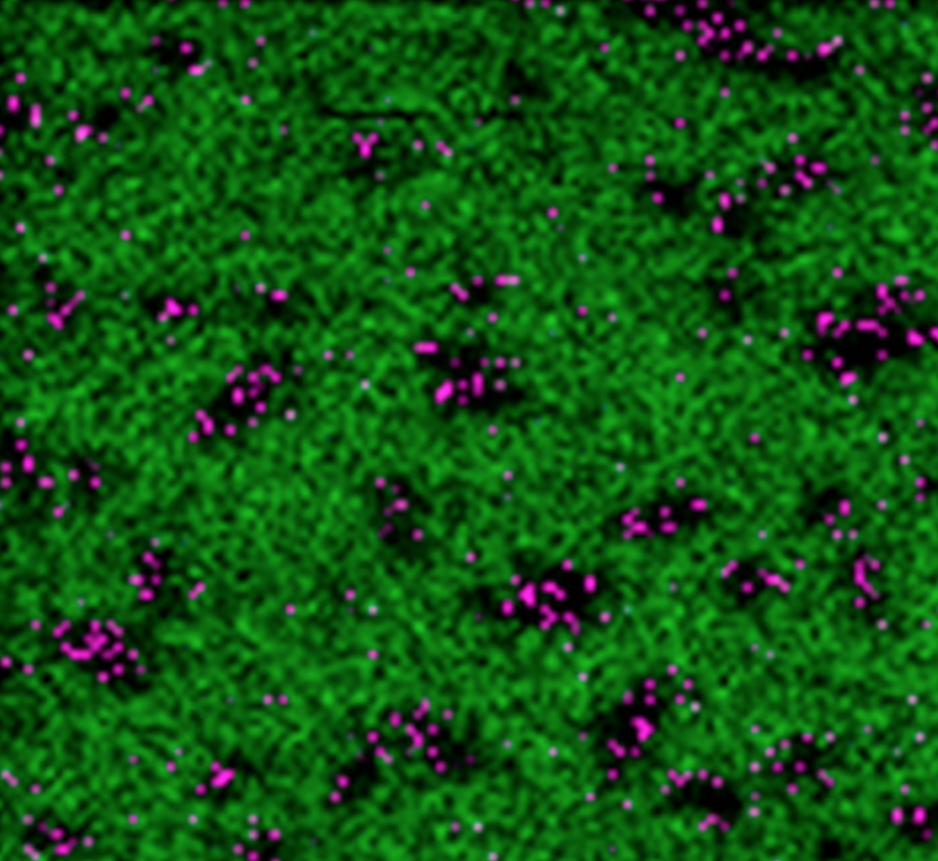 Eventually, the beams start hitting the antibiotics, which are the molecules they are trying to detect. The detection of these molecules allowed the authors to create images of the bacterial cells depicting where the antibiotics were accumulating inside of them. In one such experiment, the authors analyzed the accumulation of ampicillin inside E. coli. (See image on left.)
Eventually, the beams start hitting the antibiotics, which are the molecules they are trying to detect. The detection of these molecules allowed the authors to create images of the bacterial cells depicting where the antibiotics were accumulating inside of them. In one such experiment, the authors analyzed the accumulation of ampicillin inside E. coli. (See image on left.)
As shown, black spots (corresponding to bacteria) were speckled with purple spots (ampicillin). (The green background represents silicon, since the cells were placed on silicon wafers.)
While innovative, the trouble with the technique is that, due to random cellular fluctuations, it may not be reliably quantitative. Furthermore, it is technologically advanced, which means relatively few labs would be capable of conducting this analysis as a matter of routine. Still, in the war on antibiotic resistance, every new tool in the tool belt is welcome.
Source: Hua Tian, et al. "Subcellular Chemical Imaging of Antibiotics in Single Bacteria Using C60-Secondary Ion Mass Spectrometry." Anal Chem. Published online: 23-March-2017. DOI: 10.1021/acs.analchem.7b00466