When a patient enters a hospital or doctor's office with a cough, difficulty breathing, and chest discomfort/pain - physicians may be able to easily diagnose a lung infection. But, what is causing the infection is a different story. In fact, a physician may not be able to know - so, he or she is left to make their best guess. In this case, an antibiotic will most likely be prescribed (which is only effective against bacteria) regardless of what the cause of the infection really is.
That may not seem like a bad idea, but, prescribing an antibiotic for a viral infection is not only unnecessary - the over-prescription of antibiotics is one of the leading causes of antibiotic resistance. We know that many more antibiotic prescriptions are given out than are needed.
A new paper, published in Scientific Reports, describes a way to help physicians take the guessing game out of this decision - which would ultimately reduce the number of needless antibiotics that are prescribed by wrong guesses.
The article, entitled Transcriptomic Biomarkers to Discriminate Bacterial from Nonbacterial Infection in Adults Hospitalized with Respiratory Illness, describes the work done by the research team at the University of Rochester Medical Center (University's National Institutes of Health-funded Respiratory Pathogens Research Center).
But, the tool does not detect the invading pathogen. Rather, the idea is to analyze the body's response to the pathogen in order to identify what the pathogen is. Our immune system reacts differently to a bacterial infection than a viral infection. So, by looking at the body's response, we can determine what it is responding to - without guessing.
There were 94 adults studied, all hospitalized with lower respiratory tract infections. The team determined which patients had a bacterial infection (41 patients) and which had a viral infection (53 patients). They then identified eleven markers in the blood that could identify a bacterial infection.
This can be seen in the figure below, called a heat map. Each column represents one patient. Each row denotes a gene expressed during infection - the color indicating its level of expression. The more each gene is expressed during infection, the redder it appears on the heat map. Genes that are not highly expressed appear blue.It is easy to see the stark difference in colors between the gene expression in patients who have viral infections (clustered on the left) and those who have bacterial infections (clustered on the right.) The same genes that are blue during the viral infections (not expressed) are red in the bacterial infections (highly expressed.) It is these genes that are the basis for the tool that is under development to aid physicians in distinguishing between the two infections.
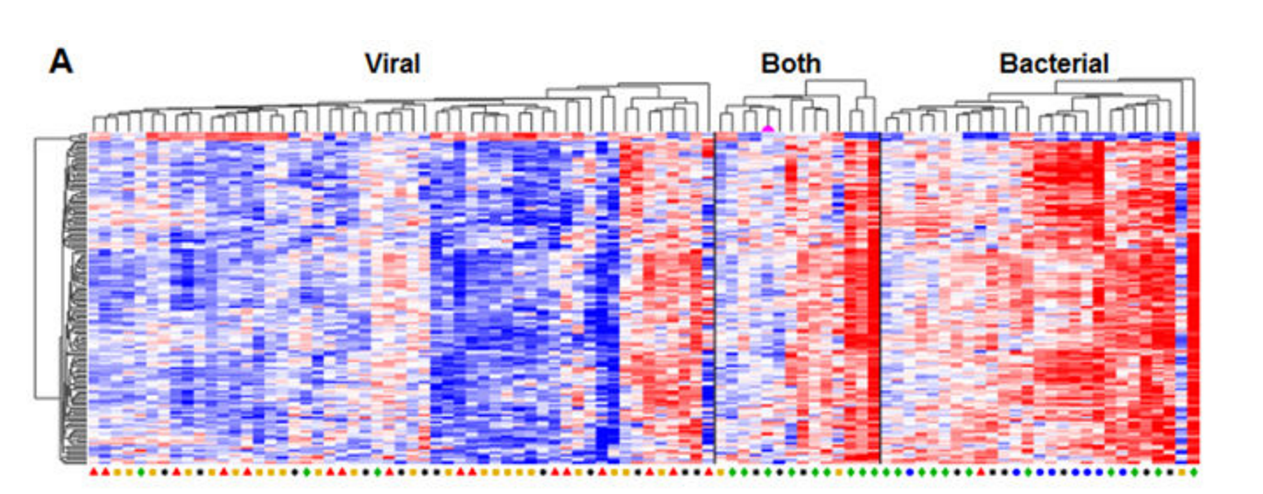
"It's extremely difficult to interpret what's causing a respiratory tract infection, especially in very ill patients who come to the hospital with a high fever, cough, shortness of breath and other concerning symptoms," said Ann R. Falsey, M.D., lead author of the study.
If this new study could lead to a tool that is accurate - and in the study, it was accurate 80 - 90% of the time - physicians may be comfortable enough to skip the antibiotic when the tool indicates a viral infection.
It looks like the project may just get the support it needs. Dr. Falsey is one of ten semifinalists in an NIH sponsored competition called the Antimicrobial Resistance Diagnostic Challenge. With an award of $50,000 to develop a prototype diagnostic test - a blood test to determine the type of pathogen causing an infection may be found in the future, in hospitals around the country.
References:
Transcriptomic Biomarkers to Discriminate Bacterial from Nonbacterial Infection in Adults Hospitalized with Respiratory Illness Bhattacharya, S. et al. Scientific Reports 7, Article number: 6548 (2017) doi: 10.1038/s41598-017-06738-3




