I have been writing for years (1) about the emergence and spread of bacteria that are resistant to virtually all antibiotics, and that it was only a matter of time until they reached the US. Time's up. They are here.
Perhaps the scariest of the bunch is carbapenem-resistant Enterobacteriaceae (CRE), which will kill about half the people who contract it. The chemical structures of amoxicillin, a member of the penicillin class and imipenem. a member of the carbapenem class are shown in Figure 1. Both classes are naturally-occurring beta-lactam antibiotics.
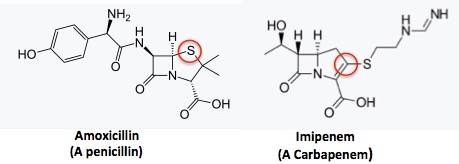
Figure 1. (Left) Amoxicillin, a member of the penicillin family. (Right) Imipenem, a member of the carbapenem family. Note the difference in the five-membered rings. Penicillins have a thiazolidine ring, which contains nitrogen and sulfur. In the five-membered ring in carbapenems, the sulfur is replaced by carbon (red circles). Five-membered rings with one nitrogen are called pyrrolidines.
Rather than rehash the news I thought I'd show how bacterial resistance is determined. It's surprisingly straightforward.
1. BACTERIAL SUSCEPTIBILITY: THE SENSITIVITY TEST
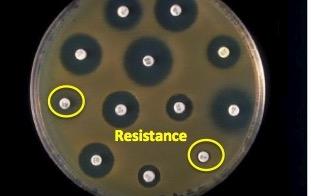
The science behind bacterial susceptibility is rather simple and logical, but also life and death. It is critical for infectious disease physicians to be able to determine what antibiotic (2) to give to a patient since an antibiotic that does not kill a bug in a Petri dish is not going to kill it inside your body. Figure 2 shows what a sensitivity test (aka the Kriby-Bauer test) looks like.
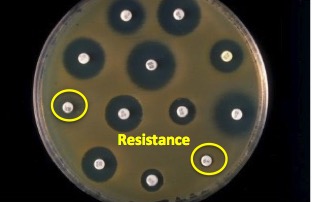
Figure 2. A bacterial sensitivity test. Image modified from Todar's Online Book of Bacteriology.
The test is done by adding bacteria to a plate which contains a growth medium (agar) and the placing paper discs that have been soaked in different antibiotics on top. The bacteria will not grow near the discs that contain an antibiotic which it is susceptible to. In Figure 2, the black circles, which are called zones of inhibition, are areas where bacteria won't grow (3). The two discs shown in yellow circles have no zone of inhibition, which means that the antibiotic on these discs will not stop this particular bug.
2. QUANTIFYING EFFECTIVENESS OF ANTIBIOTICS - THE MIC
But sensitivity is only part of the equation. Note the variation in the size of the zones of inhibition. Clearly, some antibiotics are better than others at inhibiting the growth of this particular bacterium. This parameter is expressed mathematically as the minimum inhibitory concentration (MIC). The MIC50 is the lowest concentration of the drug that will reduce bacterial growth by 50%.
The MIC provides an indirect measure of how much antibiotic will be needed to treat an infection. When the MIC value is low, the antibiotic will be very effective. When the MIC is high, it will be ineffective. Table 1 gives a striking example of the contrast between resistant and susceptible strains of the same bug - staph.
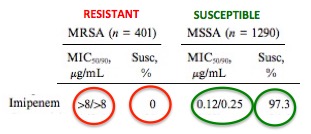

Table 1. Staphylococcus aureus : Imipenem-resistant and -susceptible strains.
The difference in efficacy of imipenem against methicillin-resistant Staphylococcus aureus (MRSA, red, left) and methicillin-susceptible Staphylococcus aureus (MSSA, green, right) is night and day. The MIC values of imipenem against MSSA range from 0.12-0.25 micrograms per mL (1290 samples), which indicates imipenem is a very potent antibiotic for treating MSSA. This is also reflected by the percentage of samples of MSSA that are killed by the drug (97.3%), so the drug will be effective against routine staph infections. But on the left (red circles), the real meaning of resistance is driven home. Even at a very high concentration (greater than 8 micrograms per mL) imipenem will not kill MRSA. Of the 401 MRSA samples tested none were killed by the drug. Source: ResearchGate
If MRSA is a problem, resistant Klebsiella pneumoniae, a bacteria that causes pneumonia and sepsis, is worse. Klebsiella has routinely been treated with carbapenems, but strains of carbapenem-resistant Enterobacteriaceae (CRE) have now been detected in the US. Although MRSA is a common and scary problem, carbapenem-resistant Klebsiella is much less common but far scarier. While there are still a number of antibiotics that can kill MRSA, Klebsiella is a much tougher cookie. If carbapenems don't work there are few alternatives. The difference in susceptible and resistant strains of Klebsiella are shown in Table 2.
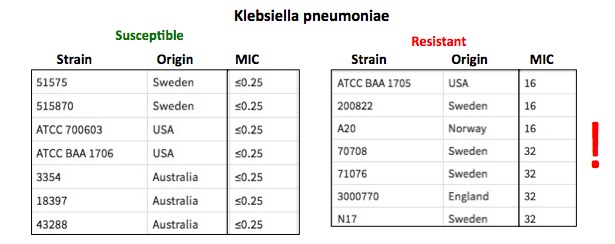
Table 2. Klebsiella pneumoniae. Meropenem-resistant and -susceptible strains.
Klebsiella strains that are sensitive to carbapenems are shown on the left and those that are resistant on the right. Note that sensitive strains are easily killed (low MIC values) by meropenem (4) while resistant strains are not. Should you be unfortunate enough to become infected by resistant Klebsiella, the options for treating the infection are dwindling. Table adapted from Nature.
3. A LITTLE PROGRESS, BUT NOT ENOUGH
In 2017 departments of health working with the CDC identified 221 cases of carbapenem-resistant enterobacteriaceae in 27 states which contained the gene that produces Klebsiella pneumonia carbapenemase (KPC), the enzyme which chemically destroys the carbapenem.
American Council advisor Dr. David Shlaes, an expert in microbiology and infectious disease research had this to say (emphasis mine):
"Most carbapenem-resistant Enterobacteriaceae (CRE) in the US carry an enzyme, KPC (originally for Klebsiella pneumonia carbapenemase), that hydrolyzes carbapenem antibiotics before they can reach their targets on bacterial cells and kill them. The enzyme was first discovered in North Carolina in 2001 and has been spreading worldwide since then. The major question is - is this due to our efforts at control as suggested by the CDC or is it because of something else like the fitness of resistant strains for example. Only time will tell. . . .
While these developments are disturbing, Dr. Shlaes also brings us a bit of good news:
This week, the CDC published a progress report entitiles Vital Signs: Containment of Novel Multidrug-Resistant Organisms and Resistance Mechanisms — United States, 2006–2017 on the numbers of CRE in the US. The data show that today, only about 4% of common Gram negative pathogens are resistant to the carbapenems. This represents almost a three-fold decrease since the peak observed in 2007 . While no one wants to have an infection with one of these pathogens, the decreasing incidence over the last decade or so is encouraging."
David Shlaes, M.D. April 4, 2018
While press coverage is dominated by silly scares like killer rubber duckies and minuscule quantities of this or that chemical in food, this is a legitimate, albeit scary, concern. The need to discover new antibiotics has never been clearer. We really need to pay attention.
NOTES:
(1) See The coming gonorrhea epidemic New York Post, 2012 and A Welcome Boost in the Race for New Antibiotics, Wall Street Journal, 2013
(2) This doesn't always apply. It takes about 1-2 days for bacteria to grow. When a patient is hospitalized with a serious infection it is impossible to wait for the results, so a "kill 'em all" antibiotic like IV vancomycin is used instead.
(3) Certain antibiotics, e.g., penicillins are bactericidal; they kill the bugs. Others, such as tetracyclines are bacteriostatic; they stop the bugs from growing but do not kill them. Either drug will be effective against a sensitive strain.
(4) Meropenem, a newer version of imipenem, is also a carbapenem.